Immune Microenvironment Analysis Group
Group Leader: Yuta Myojin
Concept: Interaction between cells/multi-omics analysis
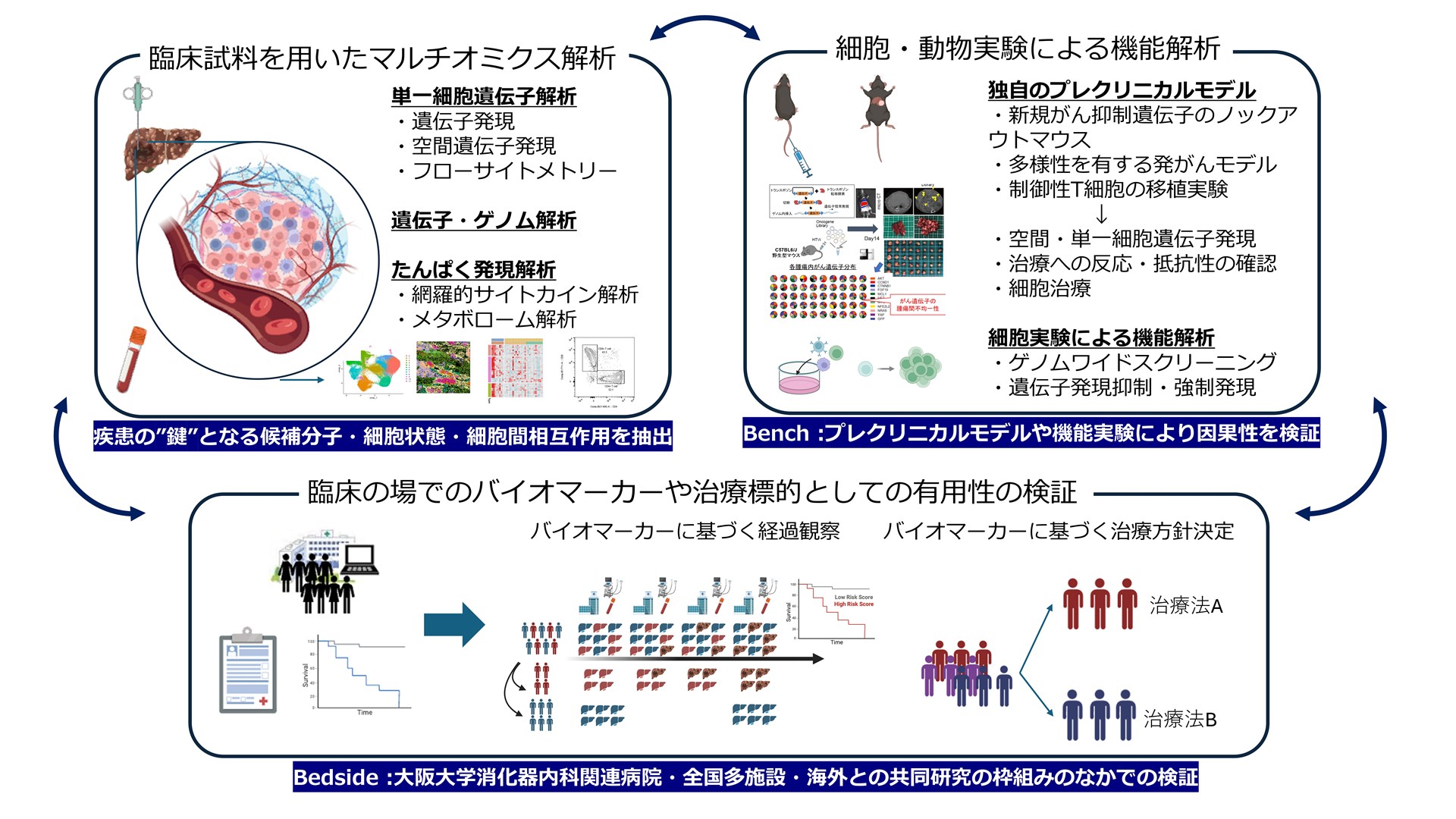
Our research group is analyzing the mechanisms by which genetic alterations in parenchymal cells, such as cancer cells and hepatocytes, and their interactions with the surrounding microenvironment, such as immune cells, fibrosis-related cells, and vascular endothelial cells, determine the establishment and progression of disease states and the response and adverse effects of drug therapies. The liver is an organ that develops and exacerbates diseases based on various background pathologies such as chronic inflammation, fibrosis, and metabolic abnormalities, and the driving factors of pathologies differ from patient to patient, even with the same diagnosis. Therefore, conventional clinical indices alone often do not fully explain disease course, prognosis, and treatment efficacy. To address this issue, our research group is integrating multi-omics analysis using clinical samples obtained from patients and various preclinical models established independently to identify key molecular mechanisms and cell-cell interactions in disease states, and to develop therapeutic targets, prognostic biomarkers, and predictors of efficacy and side effects of drug therapy. We aim to develop the technology to create therapeutic targets, prognostic biomarkers, and predict the efficacy and side effects of drug therapies. Our goal is not only to publish our research results but also to translate them into a form that can be used in clinical practice (biomarker search that contributes to the selection of treatment methods and bridges to the development of new therapeutic targets for pharmaceuticals).
Biomarker Search for Treatment Selection of Drug Therapy for Liver Cancer
Hepatocellular carcinoma has marked intra- and inter-tumor heterogeneity, and response to the same stage and treatment varies widely. In recent years, the choice of treatment has become more complex with the increasing number of immunotherapy and molecular-targeted drug options. Against this background, there is an urgent need to develop indices to determine in advance which treatment is most effective and for whom it is most likely to cause side effects. We are performing single-cell, spatial, and multi-omics analyses on a large scale using liver cancer tissue and blood samples to capture the genetic status of tumor cells and the composition and activation state of immune and stromal cells and networks of cell-cell interactions at high resolution. In addition, our uniquely established hepatocellular carcinoma model, which has heterogeneity within the same individual and more closely mimics the conditions of hepatocellular carcinoma (press release;link) to search for factors associated with the efficacy of the therapeutic agent before it is administered to the patient and the actual therapeutic agent. Through these studies, we identify candidate biomarkers by extracting cellular states that define treatment response and prognosis, as well as microenvironmental features related to treatment resistance. Our goal is to externally validate the candidate biomarkers through collaborative research with Osaka University affiliated hospitals and multiple institutions across the country, and to incorporate them into a form that can be used in medical practice.
Elucidation of the mechanism of hepatocarcinoma progression using mouse models and clinical samples
The development and progression of hepatocarcinoma cannot be explained solely by genetic alterations on the tumor cell side, but is also greatly influenced by background liver pathologies such as chronic inflammation, fibrosis, and metabolic stress, as well as dynamic changes in the tumor microenvironment. Using several mouse models, including carcinogenesis models, we are analyzing how genetic changes in cancer cells/hepatocytes alter the behavior of surrounding immune and stromal cells and drive tumor establishment and progression. For the association between hepatocyte genetic mutations and carcinogenesis, we functionally examine the pathways leading to tumor growth, immune evasion, and fibrosis promotion using knockout mice for novel tumor suppressor candidate genes identified by genetic screening in mice. We have also examined the interaction of cancer cells with hepatic stellate cells present in tumors (press release;linkThe research will be further expanded upon by the group's analysis of clinical samples to elucidate what kind of tumor microenvironment is formed in tumors and how it is involved in cancer progression, and to reconsider the phenomena of survival advantage acquisition and elimination/replacement between normal hepatocytes and precancerous/ cancerous cells in terms of genetic changes and microenvironment, and to elucidate the mechanisms leading to tumor rise We aim to elucidate the mechanisms that lead to tumor emergence, resistance to therapy, and recurrence. We believe that these studies will lead directly to the identification of pathways that drive pathogenesis as well as to the search for novel candidate therapeutic targets.
Understanding and Stratifying the Immune Microenvironment in Chronic Liver Disease
In chronic liver diseases, including autoimmune liver diseases and fatty liver, not only quantitative changes in cells constituting the microenvironment, but also changes in functional status, cell-cell interactions, and local environment define disease progression and therapeutic response. We use clinical specimens to evaluate the composition, activation state, and differentiation of immune and non-immune cells in a multilayered manner to identify immune profiles that can explain disease progression, relapse, and therapeutic response. In particular, since the microenvironment in liver tissue often exhibits different characteristics from that of peripheral blood, we will combine analyses reflecting the tissue microenvironment as needed to achieve a field-based understanding of the pathophysiology. The mechanisms that are thought to be involved in the pathogenesis identified here will be verified in animal and cell experiments, and eventually be developed into the establishment of biomarkers that will contribute to patient stratification based on immune status, optimization of treatment intensity, and prediction of relapse risk.
Future Prospects: Omics x Clinical Realization of Precision Medicine for Liver Disease
In the field of liver diseases, the need for personalized medicine is further increasing due to the growing number of treatment options, including immunotherapy, and the diversity of background liver pathologies. We aim to contribute to improving the quality of medical care for liver diseases not only by identifying the key molecular mechanisms of diseases, but also by translating them into a form that can be used in clinical practice, creating candidate therapeutic targets, establishing biomarkers for predicting prognosis, therapeutic efficacy, and side effects, and developing predictive models by integrating clinical data and omics information. We aim to contribute to the improvement of the quality of medical care for liver diseases through the development of prediction models by integrating clinical data and omics information.
Major Papers
- Myojin Y, Kodama T, Takahashi R, Nagasawa H, Kondo Y, Yusa K, Yoshida-Hashidate T, Shindou H, Furuta K, Murai K, Saito Y, Hikita H, Takehara T. Genome-wide CRISPR screen identifies ACSL3 as a regulator of lipotoxicity and progression of MASLD.
- Yamamoto S, Kodama T, Kuwano A, Maesaka K, Yoshida-Hashidate T, Shindou H, Takeda H, Shirai K, Myojin Y, Murai K, Makino Y, Tahata Y, Saito Y, Hosui A, Nozaki Y, Nakabori T, Ohkawa K, Tanaka S, Nishio A, Miyazaki M, Hikita H, Motomura K, Lujambio A, Taketomi A, Eguchi H, Takehara T. NRF2-COX2-PGE2 axis drives immune cold tumors and predicts resistance to combination immunotherapy in hepatocellular carcinoma. Hepatology. 2026 Jan 15.
- Myojin Y, Hikita H, Tahata Y, Fukumoto K, Kato S, Sasaki Y, Kumazaki S, Tsuchiya A, Enomoto M, Miki D, Yatsuhashi H, Kuroda H, Uchida Y, Yoshiji H, Yamashita T Mawatari S, Tamaki N, Miyaaki H, Asahina Y, Suda G, Matsuura K, Nakamoto Y, Hiasa Y, Takami T, Shirai K, Maesaka K, Murai K, Makino Y, Saito Y, Kodama T, Tatsumi T, Takehara T, Tatsumi Y, Tatsumi T, Takehara Y, Tatsumi Y, Kuroda H, Kuroda H, Uchida Y, Yoshiji H, Yamashita T Tatsumi T, Takehara T. Serum Growth Differentiation Factor 15 Can Be a Novel Biomarker to Predict the Prognosis of Patients With Hepatitis C Virus Hepatol Res. 2026 Jan;56(1):21-32.
- Benmebarek MR, Oguz C, Seifert M, Ruf B, Myojin Y, Bauer KC, Huang P, Ma C, Villamor-Payà M, Rodriguez-Matos F, Soliman M, Trehan R, Monge C, Xie C, Kleiner DE Wood BJ, Levy EB, Budhu A, Kedei N, Mayer CT, Wang XW, Lack J, Telford W, Korangy F, Greten TF. Anti-vascular endothelial growth factor treatment potentiates immune checkpoint blockade through a BAFF- and IL-12-dependent reprogramming of the TME. Immunity. 2025 Apr 8;58(4):926-945. e10.
- Myojin Y, Babaei S, Trehan R, Hoffman C, Kedei N, Ruf B, Benmebarek MR, Bauer KC, Huang P, Ma C, Monge C, Xie C, Hrones D, Duffy AG, Armstrong P, Kocheise L Desmond F, Buchalter J, Galligan M, Cantwell C, Ryan R, McCann J, Bourke M, Mac Nicholas R, McDermott R, Awosika J, Cam M, Krebs R, Budhu A, Revsine M, Figg WD, Kleiner DE, Redd B, Wood BJ, Wang XW, Korangy F, Claassen M, Greten TF. Multiomics analysis of immune correlatives in hepatocellular carcinoma patients Gut. 2025 May 7;74(6):983-995.
- Kumazaki S, Hikita H, Tahata Y, Sung JH, Fukumoto K, Myojin Y, Sakane S, Murai K, Sasaki Y, Shirai K, Saito Y, Kodama T, Kakita N, Takahashi H, Toyoda H, Suda G, Morii E, Kojima T, Ebihara T, Shimizu K, Sasaki Y, Tatsumi T, Takehara T. Morii E, Kojima T, Ebihara T, Shimizu K, Sasaki Y, Tatsumi T, Takehara T. Serum growth differentiation factor 15 is a novel biomarker with high predictive Aliment Pharmacol Ther. 2024 Aug;60(3):327-339.
- Monge C, Xie C, Myojin Y, Coffman-D'Annibale KL, Hrones D, Brar G, Wang S, Budhu A, Figg WD, Cam M, Finney R, Levy EB, Kleiner DE, Steinberg SM, Steinberg SM Wang XW, Redd B, Wood BJ, Greten TF. Combined immune checkpoint inhibition with durvalumab and tremelimumab with and without radiofrequency ablation in Cancer Med. 2024 Feb;13(3):e6912.
- Coffman-D'Annibale K, Myojin Y, Monge C, Xie C, Hrones DM, Wood BJ, Levy EB, Kleiner D, Figg WD, Steinberg SM, Redd B, Greten TF. VB-111 ( ofranergene obadenovec) in combination with nivolumab in patients with microsatellite stable colorectal liver metastases: a single center, single J Immunother Cancer. 2024 Jan 6;12(1):e008079.
- Myojin Y, McCallen JD, Ma C, Bauer KC, Ruf B, Benmebarek MR, Green BL, Wabitsch S, McVey JC, Fu C, Xie C, Greten TF. Adenosine A2a receptor inhibition increases the anti-tumor efficacy of anti PD1 treatment in murine hepatobiliary cancers. JHEP Rep. 2023 Nov 3;6(1):100959.
- Green BL, Myojin Y, Ma C, Ruf B, Ma L, Zhang Q, Rosato U, Qi J, Revsine M, Wabitsch S, Bauer K, Benmebarek MR, McCallen J, Nur A, Wang X, Sehra V, Gupta R, Claassen M, Wang XW, Korangy F, Greten TF. Immunosuppressive CD29+ Treg accumulation in the liver in mice on checkpoint inhibitor therapy. 509-520.
- Ruf B, Bruhns M, Babaei S, Kedei N, Ma L, Revsine M, Benmebarek MR, Ma C, Heinrich B, Subramanyam V, Qi J, Wabitsch S, Green BL, Bauer KC, Myojin Y, Greten LT McCallen JD, Huang P, Trehan R, Wang X, Nur A, Murphy Soika DQ, Pouzolles M, Evans CN, Chari R, Kleiner DE, Telford W, Dadkhah K, Ruchinskas A, Stovroff MK Kang J, Oza K, Ruchirawat M, Kroemer A, Wang XW, Claassen M, Korangy F, Greten TF. Tumor-associated macrophages trigger MAIT cell dysfunction at the HCC Cell. 2023 Aug 17;186(17):3686-3705. e32.
- Monge C, Xie C, Myojin Y, Coffman K, Hrones DM, Wang S, Hernandez JM, Wood BJ, Levy EB, Juburi I, Hewitt SM, Kleiner DE, Steinberg SM, Figg WD, Redd B, Homan P,. Cam M, Ruf B, Duffy AG, Greten TF. Phase I/II study of PexaVec in combination with immune checkpoint inhibition in refractory metastatic colorectal J Immunother Cancer. 2023 Feb;11(2):e005640.
- Matsumae T, Kodama T, Tahata Y, Myojin Y, Doi A, Nishio A, Yamada R, Nozaki Y, Oshita M, Hiramatsu N, Morishita N, Ohkawa K, Hijioka T, Sakakibara M, Doi Y, Kakita N, Yakushijin T, Sakamori R, Hikita H, Tatsumi T, Takehara T. Kakita N, Yakushijin T, Sakamori R, Hikita H, Tatsumi T, Takehara T. Thrombospondin-2 as a Predictive Biomarker for Hepatocellular Carcinoma after Cancer (Basel). 2023 Jan 11;15(2);15(2):159-160. 2023 Jan 11;15(2):463.
- Matsumae T, Kodama T, Myojin Y, Maesaka K, Sakamori R, Takuwa A, Oku K, Motooka D, Sawai Y, Oshita M, Nakabori T, Ohkawa K, Miyazaki M, Tanaka S, Mita E, Tawara S, Yakushijin T, Nozaki Y, Hagiwara H, Tahata Y, Yamada R, Hikita H, Tatsumi T, Takehara T. Circulating Cell-Free DNA Profiling Predicts the Therapeutic Cancer (Basel). 2022 Jul 11;14(14):14(14):14. 2022 Jul 11;14(14):3367.
- Myojin Y, Kodama T, Sakamori R, Maesaka K, Matsumae T, Sawai Y, Imai Y, Ohkawa K, Miyazaki M, Tanaka S, Mita E, Tawara S, Yakushijin T, Nozaki Y, Hagiwara H, Takehara T. Interleukin-6 Is a Circulating Prognostic Biomarker for Hepatocellular Carcinoma Patients Tahata Y, Yamada R, Hikita H, Tatsumi T, Takehara T. Interleukin-6 Is a Circulating Prognostic Biomarker for Hepatocellular Carcinoma Patients Treated with Combined Immunotherapy. Cancers (Basel). 2022 Feb 10;14(4);14(4)(b). 2022 Feb 10;14(4):883.
- Heinrich B, Ruf B, Subramanyam V, Myojin Y, Lai CW, Craig AJ, Fu J, Xie C, Kroemer A, Greten TF, Korangy F. Checkpoint Inhibitors Modulate Plasticity of Front Immunol. 2022 Jun 27;13:849958.
- Myojin Y, Hikita H, Tahata Y, Doi A, Kato S, Sasaki Y, Shirai K, Sakane S, Yamada R, Kodama T, Hagiwara H, Imai Y, Hiramatsu N, Tamura S, Yamamoto K, Ohkawa K, Hijioka T, Fukui H, Doi Y, Yamada Y, Yakushijin T, Mita E, Sakamori R, Tatsumi T, Takehara T. Hijioka T, Fukui H, Doi Y, Yamada Y, Yakushijin T, Mita E, Sakamori R, Tatsumi T, Takehara T. Serum growth differentiation factor 15 predicts Aliment Pharmacol Ther. 2022 Feb;55(4):422-433.
- Myojin Y, Hikita H, Sugiyama M, Sasaki Y, Fukumoto K, Sakane S, Makino Y, Takemura N, Yamada R, Shigekawa M, Kodama T, Sakamori R, Kobayashi S, Tatsumi T, Suemizu H, Eguchi H, Kokudo N, Mizokami M, Takehara T. Suemizu H, Eguchi H, Kokudo N, Mizokami M, Takehara T. Hepatic Stellate Cells in Hepatocellular Carcinoma Promote Tumor Growth Via Growth Differentiation Factor 15 Production. Gastroenterology. 2021 Apr;160(5):1741-1754.e16.
- Myojin Y, Kodama T, Maesaka K, Motooka D, Sato Y, Tanaka S, Abe Y, Ohkawa K, Mita E, Hayashi Y, Hikita H, Sakamori R, Tatsumi T, Taguchi A, Eguchi H, Takehara T. ST6GAL1 Is a Novel Serum Biomarker for Lenvatinib-Susceptible FGF19-Driven Hepatocellular Carcinoma. Clin Cancer Res. 2021 Feb 15;27(4):1150-1161.
- Suemura S, Kodama T, Myojin Y, Yamada R, Shigekawa M, Hikita H, Sakamori R, Tatsumi T, Takehara T. CRISPR Loss-of-Function Screen Identifies the Hippo Signaling Pathway as a Mediator of Regorafenib Efficacy in Hepatocellular Carcinoma. Cancers (Basel). 2019 Sep 13;11(9):1362.
